Document Type : Original Article
Authors
1 bn Sina University of Medical and Pharmaceutical Sciences, Baghdad, Iraq
2 Tikrit University, College of Sciences-Dept. Of Biology, Tikrit, Iraq
3 Baghdad-Al-Karkh Health Directorate, Al-Forat General Hospital, Baghdad, Iraq
Abstract
Background: We dealt with Multidrug- resistant Acinetobacter baumanniiisa “red alert” pathogen that is included in the acronym “ESKAPE” along with other five dangerous bacteria. The moniker “Iraqibacter” was given to this resistant bacterium isolated from American soldiers in Iraq during 2003. This is a surveillance study intended mainly to detect the antibiogram profile of this bacterium in Iraqi patients with urinary tract infection (UTI) attending a major urology center.
Materials and Methods: Two hundred ninety urine samples with positive culture results were examined for the detection of A. baumannii in a hospital-based cross-sectional studyfrom January to the end of June 2021. Isolation and identification of the bacteria, the prevalence of this bacterial infection, antibiotic sensitivity, minimum inhibitory concentration (MIC), and extended-spectrum β- lactamase (ESBL) enzyme detection were performed using conventional methods along with Vitek 2- compact 15 system (Biomerieux®).
Results: A. baumannii was isolated in 13 out of 290 UTI cases (about 4.5 %). Total resistance to ceftriaxone and cefazolin was noticed (100%), with high sensitivity to colistin and minocycline (100% and 92.3%, respectively). Nine cases of Iraqibacter isolates (about 77%) exhibited the presence of ESBL enzyme. The highest MIC value was for Trimethoprim/ sulphamethoxazole (i.e. 176.92 ± 114.56), while the least MIC detection was for colistin (0.384 ± 0.129).
Conclusion: Iraqibacter represents 4.5% of bacterial strains in UTI. Most have multidrug-resistantbehavior, with a high resistance rate to trimethoprim/ sulphamethoxazole and B-lactams, that should not be used when physicians prescribe antimicrobials to manage UTI caused by Iraqibacter.
Graphical Abstract
Keywords
Main Subjects
Introduction
In both hospital and community settings, UTI has considered the most common infection, with a global prevalence ranging between 0.7% to 19.6% in developed countries. In developing countries, as in Iraq, the prevalence is even higher (i.e., 24%) [1, 2].
Acinetobacter baumannii is a gram-negative pleomorphic bacterium that is considered an opportunistic pathogen, with a high incidence among immunocompromised individuals [3]. Since the present millennium, A. baumannii was under great concern, it is regarded as a “red alert” pathogen for both community-acquired and hospital-acquired infections due to its multidrug-resistance behavior [4-7].
Antibiotic misusage causes resistant strains to materialize in diverse illnesses, including urinary tract infections. This bacterium has garnered considerable attention in the recent decade, particularly in nosocomial settings [8].
Multidrug-resistant (MDR) bacteria are termed so because of their in vitro resistance to “more than one” antibiotic, and one of these serious bacteria that are showing this behavior is A. baumannii [9]. In the 1970s, this bacterium was sensitive to most known antibiotics, but nowadays, most A. baumannii are resistant [10,11]. The term "Iraqibacter" was used as a moniker for this resistant bacterium isolated from American soldiers in Iraq during 2003 [12].
This study aimed to report the prevalence, antibiogram profile, MIC, and ESBL enzymepresence for this MDR bacterium isolated from patients with urinary tract infections in Baghdad, Iraq.
Material and Methods
This cross-sectional hospital-based study was conducted in Al-Karamah Teaching hospital- Urology Center/ Baghdad-Iraq for six months starting from the beginning of January 2021 to the end of June 2021. We examined two-hundred ninety urine samples with positive culture results taken from Hospital-admitted patients with UTI. The detection and identification of A. baumannii were was performed using both conventional methods and an automated system for bacterial identification and susceptibility testing (i.e., Vitek 2- compact 15 system (Biomerieux ®))
Conventional methods for detection of A. baumannii: the utilized materials were ordinary and differential media (Nutrient, Blood, and MacConkey’s agar media supplied by OXOID®-England) along with several biochemical tests (Indole test, Methyl red test, Voges Proskauer test, citrate utilization test, urease test, catalase test, oxidase test, coagulase test, and nitrate test (supplied by Himedia®-India)).
The standard loop technique (one loopful carrying 0.01 ml/1µl of the urine sample supplied by Himedia®-India) was used to inoculate a drop of each urine sample into the aforementioned media to isolate different bacterial strains after overnight incubation (18-24 hours) at 37ºC in ordinary incubator (Memmert®IN55 incubator-Germany). Growing pure colonies were picked up and used for preliminary identification through their phenotypic characterization (size, color, and shape), microscopic examination (using Gram stain kit for bacterial smears-supplied by Biolab®-Hungary), and chemical properties-that are reported from the results of the used biochemical tests [13,14].
Acinetobacter baumannii bacteria are described as capsulated non-motile, strictly aerobic gram-negative coccobacilli. They are partially or non-lactose fermenters on MacConkey’s agar, catalase-positive, indole-negative, methyl-red negative, Voges Proskauer-negative, citrate-positive, oxidase-negative, coagulase- negative, urease-negative, and usually nitrate-negative [7]. They grow well on ordinary media (i.e. non-fastidious bacteria). On blood agar, they show mucoid, doomed, and non-pigmented colonies of 1 to 2 mm in diameter [15].
The A. baumannii isolates were finally identified using an automated system (i.e. Vitek 2- compact 15 system- Biomerieux®) by a fluorogenic technique for the identification of bacteria, along with a turbidimetric method for the antibiotic susceptibility testing; the antibiotic sensitivity, MIC, and ESBL enzyme were also detected using this method.
According to the manufacturer’s instructions, an applicator stick was used to transfer pure colonies in a sufficient number to 3 ml of sterile saline (pH 4.5-7.0,0.45-0.50% NaCl) placed in polystyrene test tubes. Adjustment of turbidity was performed to about 0.5-0.63 McFarland turbidity range using the DensiChek® turbidity meter (Biomerieux®). After inoculation of the identification card with the bacterial suspension, the filled card was placed into a “vacuum chamber station” in the VITEK 2 system manually. By incubating the cards (up to 30 cards) at 35.5±1.0ºC for up to 24 hours in the device, interpretation of the results using a transmittance optical system was performed [16].
Ethical Clearance was obtained from the Institutional Review Board (IRB) at Ibn Sina University of Medical and Pharmaceutical Sciences, Baghdad, IRAQ (ID: ISU.1.0.22). Patients’ informed consent was obtained from the patients for using their samples in this study.
Results and Discussion
Among 290 positive urine culture cases, only 13 (about 4.5%) of them exhibited the growth of A. baumannii in urine samples during the period of study, as presented in Figure 1.

Figure 1: The frequency of A. baumannii isolation among urine samples in current study
The antibiogram profile of A. baumannii in the current study along with ESBL detection is shown in Table 1.

Figure 2 shows the percentages of antibiotic resistance for A. baumannii. The strains isolated in the current study showed very high sensitivity to colistin and minocycline (100% and 92.3%, respectively), while total resistance to ceftriaxone and cefazolin was noticed.
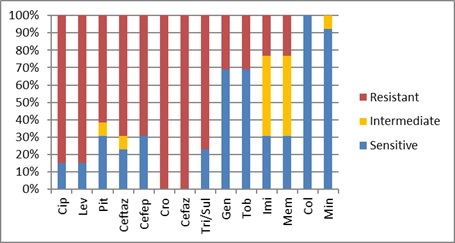
Figure 2: The frequency of antibiotic resistance for A. baumannii to the used antibiotics in current study
Figure 3 illustrates the ESBL detection frequency among A. baumannii in the current study. Nine cases of 13 (about 77%) were exhibiting ESBL positivity.

Figure 3: The frequency of ESBL detection in A. baumannii cases in current study
The mean values of MIC for these isolated A. baumanniistrains to each antibiotic are illustrated in Table 2. Trimethoprim/ sulphamethoxazole showed the highest MIC value (i.e. 176.92 ± 114.56), and the B- lactams secondly, while the least MIC detection was for colistin (0.384 ± 0.129).

Acinetobacter baumannii, a known opportunistic nosocomial microorganism, is one of the 6 most important multidrug-resistant bacteria in hospital settings worldwide and is responsible for many infections including urinary tract infection [17]. It is considered a very dangerous pathogen because of its extremely rapid ability to develop antibiotic resistance; it is now considered a resistant microorganism to almost all antimicrobial drugs that are currently present [18,19].
To our knowledge, there are few studies in Iraq having addressed the antibiotic resistance of this relatively important pathogen concerning its potential hazard [20-24]. In the current study, only about 4.5% of the total positive-culture urine samples revealed Iraqibacter isolates, which is in agreement with Radhi and Al-Charrakh [22], who found that about 5.5% of isolates from different clinical samples were Iraqibacter. Motbainor, et al., [25] also found that the frequency of MDR- A. baumanniiin different hospital-clinical isolates was about (4.6%), which also agrees with current results.
Within the last two decades, the antibiogram profile of Iraqibacter showed a progressive increase in antimicrobial resistance, which can be attributed to the link between the resistance and soil contamination of heavy metals in Iraq (i.e., the war zone), besides the already known theory of antibiotic anarchy that also led to this antimicrobial resistance [26-28]. The current study showed a multidrug-resistance pattern of the Iraqibacter to the most used drugs in our country; it is noticed, regarding Iraqi settings, that the most sensitive drugs (i.e. minocycline and colistin) are the least used ones here. This favors antibiotic misuse behavior in our country as one of the main causes of antibiotic resistance development [28].
The high prevalence of ESBL enzyme among A. baumannii isolates in the current study (69%) agrees with Singla et al. [29] who reported that up to 77% of Iraqibacter isolated from clinical samples have ESBL; it is one of the major mechanisms of this pathogen resistance to penicillin and cephalosporins, which was high in the current study. Nowadays, Iraqibacter is considered to be intrinsically resistant to these two groups of antimicrobials [30,31].
The MIC test of the fourteen antibiotics used for Iraqibacter in the current study revealed the lowest values for colistin and minocycline (0.384±0.129, 2.08±1.98, respectively) and the highest values for trimethoprim/ sulphamethoxazole (176.92±114.56). These findings agree with those of Gales et al. [29] who found similar results regarding the susceptibility pattern of Iraqibacter for these three antibiotics in the SENTRY Antimicrobial Surveillance Program (1997–2016). Kyriakidis et al. [32] also explains the high resistance rate for trimethoprim/ sulphamethoxazole, which is attributed to the presence of trimethoprim-resistant dihydrofolate reductases; in the current study, this resistance reached (77%), which is in agreement with Kanaan et al. [33], who found that the prevalence of these enzymes in A. baumannii isolates was as high as (71%) in her study.
The high values of standard deviation (SD) for MIC in current study were, perhaps, due to the presence of few isolates (i.e., No. 7, 9 and 13 in table (1)) with high susceptibility to commonly used antibiotics. This widely affects the SD of the MIC besides the low number of Iraqibacter isolates generally.
Conclusions
Multidrug-resistant A. baumannii (Iraqibacter) represents about 4.5% of bacterial pathogens in UTI, with a high resistance rate to trimethoprim/ sulphamethoxazole and B-lactams. These groups of antimicrobials should not be used by physicians who tried to manage UTI caused by Iraqibacter. The physicians need to ask for culture and susceptibility tests before prescribing antibiotics to prevent the gradual increase in MIC for bacterial strains causing UTI.
Acknowledgments
The authors provide their sincere gratitude to the laboratory workers of the Urology Center at Al-Karamah Teaching Hospital for their generous assistance in facilitating the collection of the urine samples.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Authors' contributions
All authors contributed toward data analysis, drafting and revising the paper and agreed to responsible for all the aspects of this work.
Conflict of Interest
We have no conflicts of interest to disclose.
ORCID:
Sinan Bahjat Alrifai
https://www.orcid.org/0000-0003-1585-2401
Wakas Saadi Mahmood
https://www.orcid.org/0000-0002-2737-1173
Noor Hamdi Jasem
https://www.orcid.org/0000-0002-3049-0851
HOW TO CITE THIS ARTICLE
Sinan Bahjat Alrifai, Wakas Saadi Mahmood, Noor Hamdi Jasem. Surveillance of Multidrug-resistant Iraqibacter Isolated from Patients with Urinary Tract Infection at a Baghdad Urology Center, J. Med. Chem. Sci., 2022, 5(5) 753-759
- Medina M., Castillo-Pino E., Adv. Urol., 2019, 11:175628721983217 [Crossref], [Google Scholar], [Publisher]
- Tandogdu Z., Wagenlehner F.M., Opin. Infect Dis., 2016, 29:73 [Crossref], [Google Scholar], [Publisher]
- Montefour K., Frieden J., Hurst S., Helmich C., Headley D., Martin M., Boyle, D.A., Care Nurse, 2008, 28:15 [Crossref], [Google Scholar], [Publisher]
- Manchanda V., Sinha S., Singh N., Glob. Infect. Dis., 2010, 2:291 [Crossref], [Google Scholar], [Publisher]
- Abbo A., Navon-Venezia S., Hammer-Muntz O., Krichali T., Siegman-Igra Y., Infect. Dis., 2005, 11:22 [Crossref], [Google Scholar], [Publisher]
- Cerqueira G., Peleg A., IUBMB Life, 2011, 63:1055 [Crossref], [Google Scholar], [Publisher]
- Peleg A.Y., Seifert H., Paterson D.L., Microbiol. Rev., 2008, 21:538 [Crossref], [Google Scholar], [Publisher]
- Jiménez-Guerra G., Heras-Cañas V., Gutiérrez-Soto M., del Pilar Aznarte-Padial M., Expósito-Ruiz M., Navarro-Marí J.M., Gutiérrez-Fernández, J., Med. Microbiol., 2018, 67:790 [Crossref], [Google Scholar], [Publisher]
- Magiorakos A.P., Srinivasan A., Carey R.B., Carmeli Y., Falagas M.E., Giske C.G., Monnet D.L., Microbiol. Infect., 2012, 18:268 [Crossref], [Google Scholar], [Publisher]
- Fournier P.E., Vallenet D., Barbe V., Audic S., Ogata H., Poirel L., Claverie J.M., PLoS Genet. 2006, 2:e7 [Crossref], [Google Scholar], [Publisher]
- Jiang M., Chen X., Liu S., Zhang Z., Li N., Dong C., Zhang L., Wu H., Zhao S., Public Health, 2021;9 [Crossref], [Google Scholar], [Publisher]
- MMWR Morb. Mortal. Wkly. Rep., 2004, 53:1063 [Google Scholar], [Publisher]
- Grabe M., Bjerklund-Johansen T.E., Botto H., Çek M., Naber K.G., Tenke P., Wagenlehner F., Urol., 2015, 182:237 [Google Scholar]
- Atlas R.M. Handbook of Media for Environmental Microbiology. 2nd Boca Raton: Taylor & Francis; 2005 [Crossref], [Google Scholar], [Publisher]
- Alsan M., Klompas M., Clin. Outcomes Manag., 2010, 17:363 [Google Scholar], [Publisher]
- Pincus D.H., Bethesda, MD: Parenteral Drug Association, 2006, 2:1 [Google Scholar], [Publisher]
- Antunes L., Visca P., Towner K., Dis., 2014, 71:292 [Crossref], [Google Scholar], [Publisher]
- Poirel L., Walsh T., Cuvillier V., Nordmann P., Microbiol. Infect. Dis., 2011, 70:119 [Crossref], [Google Scholar], [Publisher]
- Poirel L., Nordmann P., Microbiol. Infect., 2006, 12:826 [Crossref], [Google Scholar], [Publisher]
- Scott P., Deye G., Srinivasan A., Murray C.., Moran K., Hulten E.D., Petruccelli B., Infect. Dis., 2007, 44:1577 [Crossref], [Google Scholar], [Publisher]
- AL-Kadmy I., Ali A., Salman I., Khazaal S., New Microbes New Infect., 2018, 21:51 [Crossref], [Google Scholar], [Publisher]
- Radhi S.H., Al-Charrakh A.H., Indian J. Public Health Res. Dev., 2019, 10:668 [Crossref], [Google Scholar], [Publisher]
- Mshachal M.A., Abdulrahman T.R., Khudhair M.S., Hassan J.S., Iraqi J. Med. Sci., 2017, 15:314 [Crossref], [Google Scholar], [Publisher]
- Ibrahim S., Al-Saryi N., Al-Kadmy I., Aziz S., Mol Biol Rep., 2021, 48:6987 [Crossref], [Google Scholar], [Publisher]
- Motbainor H., Bereded F., Mulu W., BMC Infect. Dis., 2020, 20:92 [Crossref], [Google Scholar], [Publisher]
- Dewachi O., Skelton M., Nguyen V.K., Fouad F.M., Sitta G.A., Maasri Z., Giacaman R. Lancet, 2014 383:449 [Crossref], [Google Scholar], [Publisher]
- El-Sayed M.H., J. Microbiol. Res., 2016, 4:26 [Crossref], [Google Scholar], [Publisher]
- Truppa C., Abo-Shehada M.N., BMC Infect. Dis., 2020, 20:936 [Crossref], [Google Scholar], [Publisher]
- Singla P., Sikka R., Deeep A., Gagneja D., Chaudhary U., Clin. Diagn Res., 2014, 8:D16 [Crossref], [Google Scholar], [Publisher]
- Kumar S., Anwer R., Azzi A., Microorganisms. 2021, 9:2104 [Crossref], [Google Scholar], [Publisher]
- Gales A.C., Seifert H., Gur D., Castanheira M., Jones R., Sader H., Open Forum. Infect. Dis., 2019, 6:S34 [Crossref], [Google Scholar], [Publisher]
- Kyriakidis I., Vasileiou E., Pana Z., Tragiannidis A.., Pathogens. 2021, 10:373 [Crossref], [Google Scholar], [Publisher]
- Kanaan M.H.G., Al-Shadeedi S.M.J., Al-Massody A.J., Ghasemian A., Immunol. Microbiol. Infect. Dis., 2020, 70:101451 [Crossref], [Google Scholar], [Publisher]


