Document Type : Original Article
Authors
1 Al-Zahraa Teaching Hospital, Wasit Health Directorate, ministry of Health, Iraq
2 Department of Clinical Laboratory Science, College of Pharmacy, University of Baghdad, Baghdad, Iraq
Abstract
Background: Transmembrane multiligand receptor RAGE is a protein found on the surface of cells that binds to the advanced glycation end products. Platelet cyclooxygenase activity is elevated in response to RAGE activation, contributing to the formation of a thrombus.
Objectives: To examine the connection between RAGE gene rs80096349(C>T), rs1035798(C>T), and rs184003(G>T) polymorphisms and resistance to aspirin in CAD patients from Iraq.
Patients and methods: Patients with coronary artery disease (CAD) (161 males and 64 females) were enrolled in the trial between February 2021 and October 2021. All participants were taking prophylactic Aspirin (100 mg). The control group consisted of 130 individuals, including 97 males and 33 females, who appeared to be in good health and were not taking aspirin. Serum thromboxane B2 levels were used to assess the effectiveness of aspirin (TBX2). Patients were classified as either aspirin sensitive or resistant. The polymorphism of RAGE's most closely related single nucleotide polymorphisms (SNPs) was identified using polymerase chain reaction for amplification of the extracted deoxyribonucleic acid and Sanger's method of sequencing.
Results: A total of 225 cardiovascular patients compatible with inclusion criteria were enrolled. There were 161(71.6%) male and 64(28.4%) female patients with a mean age of 56.85±8.11years old. The statistical analysis revealed 17.8% (n = 40) aspirin-resistant cases in the study population. Genotypic analysis of our data showed that participants with the T allele of rs184003(G/T) had an increased prevalence in the aspirin-resistant group compared with the aspirin-sensitive group (p<0.05). In contrast, for rs10835798, the frequency was dramatically higher for T alleles in aspirin sensitive group contrasted with the resistant group (p<0.05). The aspirin-resistant group was found to have significantly greater RAGE levels than the sensitive group.
Conclusion: Our results provide evidence that RAGE expression and gene rs1035798(C>T), and rs184003(G>T) polymorphisms could be predictors for CAD patients' resistance to aspirin.
Graphical Abstract
Keywords
Main Subjects
Introduction
In terms of cardiovascular mortality, coronary artery disease (CAD), and diabetes are the top non-communicable diseases worldwide [1-4], and they have serious complications [5, 6]. Therefore, these diseases have become the target for many researchers [7]. Atherothrombosis is the major pathological alteration and primary cause of cardiovascular disease, and platelets play a crucial role in this process [8].
The aspirin use for both the primary and secondary prevention of cardiovascular disease is widespread [9]. Aspirin's capacity to permanently acetylate platelet cyclooxygenase-1 (COX-1) and hence suppress thromboxane A2 production by platelets is responsible for its therapeutic efficacy in cardiovascular disorders [10].
However, many people may not respond to aspirin despite its widespread usage for a wide variety of conditions. These people are also known as non-aspirin responders [11]. This condition, also known as aspirin resistance, has received a lot of attention from researchers. It is also connected to the idea of aspirin's diminished effectiveness in preventing cardiovascular disease (clinical resistance) and its ineffective inhibition of platelet and TXA2 production [12].
A gene that codes for advanced glycation end products receptor (RAGE) has been pinpointed to chromosome 6p21 [13]. RAGE is overexpressed in disease states such as cardiovascular disease, diabetes, and inflammation relative to healthy animal models and people [14]. It has the capability of binding several different ligands due to the specificity of its domain [15, 16]. Among many cell types that express RAGE receptors are platelets [17].
A large body of research has established that, under pathological circumstances, RAGE plays a significant role in platelet hyperactivation and thrombus formation [18, 19]. Vascular problems in patients with coronary artery disease (CAD) and diabetes mellitus (DM) have been investigated extensively, and the results reveal a correlation between RAGE gene polymorphism and both conditions [20]. There are more than 30 types of polymorphisms in the AGER gene, the vast majority of which are single nucleotide variants (SNVs) that influence RAGE gene expression and function. Knowledge of which of these gene variants is linked to the emergence of aspirin resistance is crucial for the early detection in patients with specific SNVs, the creation of quick diagnostic kits, and the future
formulation of therapy programs. Yet, to the best of our knowledge, no clinical investigation has investigated the potential link between RAGE gene polymorphisms and the development of Aspirin resistance. In this study, we predict that the RAGE gene polymorphisms rs80096349(C>T), rs1035798(C>T), and rs184003(G>T) could be correlated with aspirin resistance in Iraqi patients with coronary artery disease (CAD).
Study design
Patients and method
The Department of Cardiology at Al-Zahraa Teaching Hospital, Wasit Health Directorate, undertook a convenient cross-sectional study between February and October of 2021. The current research was a subset of a larger study that included 225 Iraqi CAD patients who met the study's inclusion criteria for convenience sampling. We began with 232 patients who had stable coronary artery disease (CAD). Only 225 people total (161 males and 64 females) were included in the analysis (excluding two patients due to noncompliance; five patients due to non-valid samples). All the patients who participated had been taking a daily 100 mg dose of Aspirin for at least seven days prior to enrollment.
Inclusion criteria
Patients should be 18 years or older, have stable coronary artery disease with or without type 2 diabetes, and be receiving treatment with oral hypoglycemic medications and/or insulin. Everyone was evaluated for their condition and given care by an expert in the field.
One of the following is required as a definition of CAD [21]:
- Coronary artery stenosis of at least 50% (at least one) as detected by cardiac catheterization.
- Previous PCI or previous MI within the past six months.
- Surgery to bypass the coronary arteries (about a year ago).
- The presence of ST depression of at least 1 mm in at least two contiguous ECG leads is considered diagnostic of an abnormal exercise treadmill test.
- Stress echocardiography or pharmacological stress followed by revascularization.
One of the following conditions should be met to diagnose type 2 diabetes: An HbA1c level equal to or more than 6.5% and/or Having a fasting blood glucose (FBG) level equal to or more than 126 mg/dL (after fasting for at least 8 hours) [22].
Exclusion criteria
- When platelet function may be altered using anticoagulants or any other medicine.
- Treatment with heparin or low-molecular-weight heparin during the preceding 24 hours of participation.
- History of family or personal bleeding disorders.
- Haemoglobin<8 g/dl.
- A myocardial infarction that is acutely occurring during the past 30 days.
- Procedures for coronary angioplasty performed within the last 30 days.
- Within a six-month time frame of a stroke happening.
- Cooperating surgical procedures, such as a valvuloplasty or a maze surgery.
- Liver disease and renal impairment.
- Insufficient or excessive platelet count, defined as either less than 150×109/l or greater than >450×109/l.
- Greater than 1.2 for the international normalized ratio.
- Chronic inflammation and neoplastic diseases.
- Female patients who were pregnant.
Patients were fully briefed on the objectives and scope of the study, and participation in the research was restricted to only those who gave their informed permission in writing.
The control group was healthy and had avoided NSAIDs and aspirin for at least ten days. Ninety-seven males and 33 women made up the 130-person group. No control individual had ever experienced a vascular incident before the experiment. The control group was needed to define the standard reference range for TBX2. College of Pharmacy at the University of Baghdad's Scientific and Ethical Committee and the Research and Development Committee of Wasit Health Directorate both gave their stamps of approval to the procedures we followed. In accordance to the tenets of the Declaration of Helsinki, this study was carried out.
Serum TBX2 level and other biochemical tests
After collecting a blood sample in an EDTA tube, it was centrifuged at 3000 rpm for 10 minutes. Serum samples were aliquoted into 1 mL Eppendorf tubes for biochemical analysis and frozen at (-80 °C) for later use. Immediate biochemical analysis was performed.
Data collection
Patients were interviewed using a data collection form to get demographic information (such as age, gender, body mass index, disease duration, and so on) and a complete medical history. Commercially available ELIZA kits (SHANGHAI YEHUA Biological Technology Co., Ltd.CHINA) have been used to conduct assays for TBX2 in thawed samples, with the manufacturer's protocol being followed [23].
Aspirin resistance diagnostic criteria
Aspirin's effect on platelet COX-1 activity is measured using serum thromboxane B2(TBX2) test [24]. The reference range of our study obtained by the control group was (407.82-671.47) pg/mL. Accordingly, if the serum TBX2 level drops to below 97% of the lower limit of the reference range (12.23 pg/mL), then the patient is regarded to be aspirin sensitive. Otherwise, the patient is considered to be aspirin resistant [25]. Next, the patients were classified as either aspirin sensitive or resistant. After identifying the group of resistant individuals, a similar number of patients from the sensitive group were selected randomly to do the necessary comparison and analysis (including polymerase chain reaction [PCR] and DNA sequencing) (Figure 1). Serum levels of aspirin were used to assess patients' adherence to the treatment plan [26], besides interviewing patients regarding aspirin adherence.

Figure 1: Distribution of subjects who participated in the current study
Primers
The lyophilized primers (Table 1) were supplied by Macrogen Company. Lyophilized primers were dissolved in nuclease-free water to a final concentration of 100 pmol/µl as a stock solution. A working solution of these primers was made by adding 10 µl of primer stock solution (stored at -20 °C) to 90l of nuclease-free water to yield 10 pmol/µl.
Amplifications of PCR and the optimization of primers
(Forward) (Reverse) primers were used to amplify the template of DNA at 55, 58, 60, 63, and 65 °C to determine the best annealing temperature. 20 µl volumes included 10 µl GoTaq Green Master Mix (2X), 1l for each primer (10 pmol), 6l nuclease-free water, and 2l of template DNA. PCR was done with PCR Express (BioRad, USA) and the following temperature program: Denatured at 94 °C for 4 min, and then 30 cycles of denaturation, annealing, and extension at 72 °C for 30 seconds. A final 7-minute incubation at 72 °C followed by 10 minutes at 4 °C to stop the reactions (Figure 2).
Sequencing of PCR products
Macrogen Corporation-Korea's ABI3730XL is an automated DNA sequencer that was used to sequence PCR products using the Sanger sequencing method. After receiving the findings through email, geneious software was used to do the analysis.

Statistical analysis
The program IBM SPSS for Windows version 26.0 was utilized to carry out the statistical analysis (IBM Corp., Armonk, NY, USA).
The values of continuous variables were reported as the mean accompanied with the standard deviation. The numerical and percentage representations of the categorical variables were given. To determine whether or not the variables in the findings followed a normal distribution, the Kolmogorov-Smirnoff test was carried out.
To establish whether there was a statistically significant difference in the demographic parameters of the sensitive group and the resistant group, an unpaired t-test was carried out on the normally distributed data. The Chi-square test was utilized to investigate the possibility of proportional differences between groups. The TBX2 reference interval was derived from the measurements of healthy people with the same or comparable features. After sorting the data, we used the nonparametric technique, which was based on determining the 2.5 and 97.5 percentiles of the distribution. The independent determinants of aspirin resistance were identified through the use of binary logistic regression analysis. A significant difference was shown for all the tests where P was less than 0.05.
Results and Disussion
Table 2 illustrates the demographic, clinical, and biochemical data of the participating patients. They were matched regarding age, gender, and BMI. While CRP, TBX2, and RAGE serum levels were markedly higher in the group that was resistant to aspirin as compared with the group that was responsive to aspirin (P<0.05). Furthermore, the other biochemical markers examined showed no statistically significant differences (P > 0.05).
Aspirin resistance frequency in the studied population
The aspirin resistance frequency of our study was (17.8%), where forty of the total patients were resistant to aspirin, as depicted in Figure 3.


The distribution of RAGE gene polymorphism by genotype and allele frequency in resistant and sensitive populations
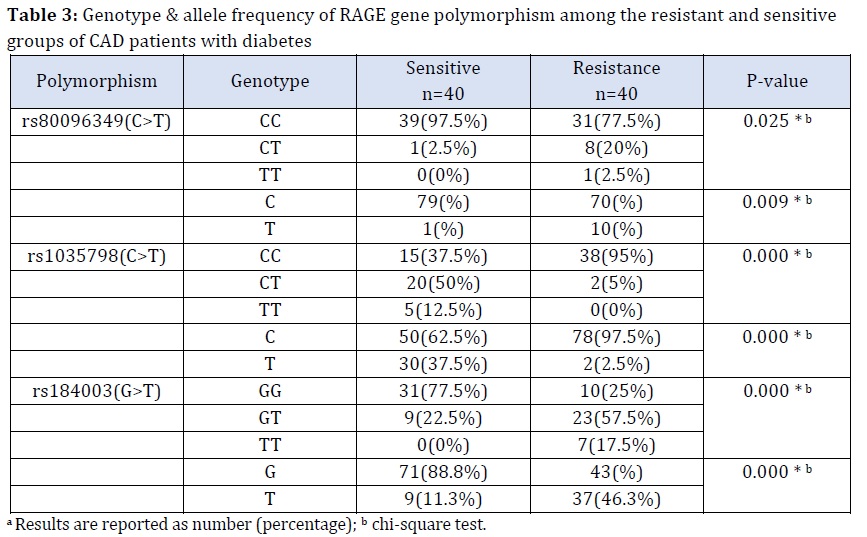
Data on the genotype and allele frequencies of the RAGE polymorphisms under study are summarized in Table 3 for both susceptible and resistant individuals. The heterozygote CT, GT genotypes, and there was a statistically significant increase in the number of resistant individuals who had the mutant homozygote TT genotype compared with the number of susceptible individuals for rs80096349 (C>T) and rs184003(G>T), respectively. Likewise, the mutant T alleles were significantly higher in the resistant groups compared with the sensitive groups for both SNPs (p<0.05), as is explained in Table 3. Noteworthy, the frequency of the wild CC genotype of rs1035798 (C>T) was significantly higher in the resistant groups compared with the sensitive groups, while the mutant CT and TT genotypes were prevalent in the sensitive compared with the resistant group. Furthermore, there was a marked increase in the frequency of the mutant T allele in the sensitive groups compared to the resistant groups (Table 3) (p<0.05).
Association of mean serum level of RAGE and other studied parameters with rs80096349 (C>T), rs1035798 (C>T), and rs184003 (G>T) RAGE gene polymorphisms of participating patients
Focusing on the association of serum RAGE level and the other studied parameters with the RAGE polymorphisms, and because of the limited number of people that carry the mutant homozygote genotype, this group was included for statistical analysis with those with the heterozygote genotype for all significant SNPs. Table 4 showed that the mutant CT+TT genotypes for rs80096349(C>T) and mutant GT+TT for rs184003(G>T) were characterized by significantly higher RAGE and CRP serum levels compared with the patient that had GG genotype (p <0.05).
Table 4 showed a significant increase in RAGE and CRP levels in the homozygote CC genotype compared with mutant heterozygotes CT+TT regarding rs1035798 (C>T). All other parameters showed non-significant differences (p >0.05).
Patient risk factors and their correlations with serum RAGE levels
Table 5 illustrates Spearman's correlation studies of RAGE with some studied risk variables. RAGE serum levels had a positive correlation which was significant with CRP serum level (r=0.51; P<0.05), presence of smoking (r=0.31; P<0.05), smoking intensity (r=0.33; P<0.05), and smoking duration (r=0.38; P=0.05%). While the remaining parameters showed either a positive or negative correlation, none of them was statistically significant (P > 0.05).


Aspirin resistance correlation with the included patient risk factors
Spearman's correlation studies of aspirin resistance with the studied variables were explained in Table 6. Aspirin resistance had a moderate positive correlation which was significant with CRP serum level (r=0.381; P<0.05), RAGE serum level (r=0.49; P<0.05), frequency of rs80096349 (C>T) (r=0.30; P<0.05), and frequency of rs184003 (C>T) (r=0.52; P=0.05%). On the other hand, a significant negative correlation between aspirin resistance and the frequency of rs1035798(C>T) (r=-0.60; P=0.000), indicating the protective role of this variant, while none of their remaining parameters showed a significant correlation (P>0.05).
Analysis of risk variables for Aspirin resistance using binary logistic regression
Aspirin resistance was studied using binary logistic regression to examine the impact of the most linked independent risk factors, as presented in Table 7.
Forward Stepwise was employed to select the important determinants of aspirin resistance among patients using SPSS Software. Accordingly, the frequency of the mutant T allele of rs184003 (G>T), the dominant (C) allele of re1035798 (C>T), and serum RAGE levels were found to be the most significant predictors of aspirin resistance (Table 7).
The model revealed that with each increase in RAGE serum level by 1ng/ml, the likelihood of the patient being aspirin resistant is increased by greater than six times (odds ratio:7.696), and the mutant T allele of rs1035798(C>T) has a protective effect where patients having the (T) allele is more than ninety-nine percent sensitive to aspirin (odds ratio:0.001) compared with patient carrying wild-type homozygote CC. Likewise, the probability of a patient carrying the T allele of rs184003(G>T) being aspirin resistant is sixteen times more likely compared with a patient with wild type (GG) of rs184003(G/T) (odds ratio:17.27).
Aspirin resistance is considered as a potential contributor to the recurrent coronary events in the populations of different ethnicity. Variations in platelet function testing procedures, definitions of aspirin resistance, and racial/ethnic background all contributed to the broad ranges of aspirin resistance reported by researchers (5-45%) [27]. Our study found that patients non-responding to aspirin were forty, represented about one-fifth of the total CAD patients (Figure 3). Aspirin resistance is multifactorial, and several contributing factors have been considered, including cigarette smoking [28], oxidative stress [29], and the most important one, the genetic polymorphism [30]. The effect of cigarette smoking on CAD and diabetes has been widely studied [31, 32]. Several studies showed that cigarette smoking has a positive correlation with aspirin resistance in different clinical settings [28]. However, another studies showed no correlation [33-35]. This variability in findings may be related to sample size, the type of studied population, and the presence of other factors.

Our findings revealed that cigarette smoking, intensity, or duration are not risk factors for aspirin resistance (Table 6). However, there was a significant correlation between cigarette smoking with RAGE serum levels (Table 5) in line with the other studies [36].
For its role as an inflammatory biomarker in cardiovascular disease, C-reactive protein (CRP) has been the subject of much research [37, 38]. The oxidative stress has been linked to the cardiovascular disease and diabetic problems in a number of studies [39-41]. The evidence between oxidative stress and thromboxane A2 production has been shown in various clinical scenarios with the elevated risk for atherothrombosis [42]. Aspirin-resistant platelet aggregation is thought to be caused by oxidative stress, which stimulates the formation of thromboxane A2 from arachidonic acid via nonenzymatic or enzymatic processes (such as via lipoxygenase) that aspirin does not inhibit [43], pointing to a possible link between the oxidative stress and aspirin resistance [44].
It is important to identify single nucleotide polymorphisms (SNPs) that may be associated with aspirin resistance to better understand why certain patients do not respond to aspirin and to create a novel antiplatelet therapeutic approach.
To our best knowledge, the association between RAGE gene polymorphisms and the incidence of aspirin resistance across all illness settings is a new concept worldwide. Hence, there is no similar study to compare our results with it. Therefore, we will give some rational reasons to explain our results as much as possible. RAGE belongs to the immunoglobulin superfamily and can interact with several ligands of a wide variety of cells, including platelets [17].
Polymorphisms in the RAGE gene may be an important negative marker of coronary disease [45]. RAGE itself could be used as an oxidative stress biomarker [46]. So, disease biomarkers may include genetic variations that impact RAGE mRNA or protein levels.
According to the previous studies, antioxidant status has been linked to certain RAGE polymorphisms [47, 48]. Persons having the T allele of rs184003 (G>T) polymorphism had compared to individuals carrying the G allele, those carrying the T allele have considerably lower plasma levels of numerous antioxidants (total carotenoids, lutein, lycopene, and tocopherol) [49].
In agreement with the above assumptions, our study revealed that serum RAGE and CRP levels in patients carrying GT+TT genotype of rs184003 and CT+TT genotypes of rs80096349(C>T) were significantly higher than in patients having homozygous GG and CC genotype, respectively (Table 4).
The intron site's functional significance is unclear in general, so the rs184003 (G>T) mutation did not affect gene translation directly. It may have indirect effects on the splicing of the RAGE mRNA. In another way, mechanisms under consideration include aberrant mRNA splicing, altered mRNA stability, and connections to neighboring coding SNPs [49, 50].
In relation to the very rare rs80096349(C>T) SNP, the mutant T variant is present in just ten (12.5%) of the total included patients (Table 3). Notably, there has been no clinical study till now evaluating its role [51]. It is entirely novel and discovered through investigation, while we are making DNA sequencing our samples, so the explanation of the exact mechanism by which this variant increases RAGE level is not mentioned till now and requires further studies.
The opposite results were found regarding rs1035798 (C>T), where serum RAGE and CRP levels in patients carrying homozygote CC genotype were significantly higher than in patients having mutant CT+TT genotype (Table 4). It has been hypothesized that the presence of the SNP's minor T allele interferes with splicing factor recognition [52], leading to RAGE upregulation. Our study reveals that the mutant T allele of rs1035798 (C>T) has a protective effect, that is in agreement with the other studies that showed that this variant has a protective effect in different settings [53].
Furthermore, our findings showed that the serum RAGE level was correlated significantly with the CRP serum level (r:0.51; p: 0.00), as is seen in Table 5. This is supportive evidence for a positive correlation between RAGE expression and oxidative stress.
Our study from the binary logistic regression model (Table 7) revealed that the mutant T allele frequencies of rs184003 (G>T), the dominant (C) allele of re1035798 (C>T), and serum RAGE levels were found to be the most significant predictors for aspirin resistance in the studied population.
Since this is a novel study, explaining the exact mechanisms by which RAGE gene polymorphisms modulate the response to aspirin requires further investigations.
We can give some rationale explanations. First, classical acute phase reactants, such as C-reactive protein, were shown to be elevated after RAGE activation by its ligands [54, 55]. On the other hand, CRP up-regulates the RAGE expression [56, 57]. Therefore, the conclusion is that the oxidative stress may be the connecting factor between RAGE expression and resistance to aspirin. Second, RAGE can directly activate platelet and cause thrombotic complications through binding to AGEs of different sources, especially in diabetics [58], S100A12 ligand [18], or HMGB1 ligand [59] and because of their synergistic effect, lowering expression of HMGB1 and RAGE might have a major impact on preventing cardiovascular problems [60]. Therefore, the therapeutic targeting of the HMGB1-RAGE pathway may be useful for the prevention and treatment of vascular damage in thrombosis disease [60]. Third, it is widely accepted that neutrophil recruitment is crucial for platelet thrombus stabilization. RAGE ligand HMGB1 is the most effective neutrophil messenger. In patients with acute myocardial infarction treated by thrombus ablation, HMGB1 has been shown to bind to neutrophil RAGE and up-regulate autophagolysosome formation via the MAPK pathway [61].
Fourth, according to a review paper published by Ajjan et al., aspirin resistance is more common in persons with diabetes because the elevated glycation of platelet and coagulation factor proteins disrupts the acetylation process. Finally, upregulation of adhesion molecules and pro-coagulant activity, decreased platelet survival, increased platelet aggregation, and altered levels of coagulation factors like anti-thrombin 3, tissue factor, thrombomodulin, and fibrinolytic inhibitor plasminogen-activator inhibitor-1 are all results of AGE-induced activation of RAGE.
Limitations
This study has many limitations. First: the sample size is modest, which may be due to the difficulty in recruiting patients who meet the study's eligibility requirements. Second, the study was done in a single center. Therefore, it would be unwise to extrapolate our results to people of different races without further investigation. Third, currently, only the three most often investigated polymorphisms of RAGE have been genotyped, but future research can expand this to include the other functional polymorphisms. Fourth, the study lacks patient follow-up to evaluate the patient clinically concerning the incidence of thrombotic complications especially in aspirin resistant patients due to the lack of time for conducting a cohort study. Finally, the insufficient financial funds were another limitation in this study.
Conclusion
Analysis of Iraqi patients revealed a rather high prevalence of aspirin resistance. The experts who are thinking about prescribing aspirin to patients with CAD should be aware of this discovery, since it may pose a concern for them. This study demonstrates evidence that the examined RAGE gene variants are linked to an increased likelihood of aspirin resistance. Thus, these SNPs have the potential to serve as a diagnostic biomarker to estimate the number of people at high risk for CAD who are resistant to aspirin.
Acknowledgments
The authors appreciate the efforts of everyone who contributed to the completion of this work.
Funding
The authors of the study did not receive any financial support for the research.
Authors' contributions
Each author has acknowledged the full responsibility for all elements of this work and has made substantial contributions to its development (through analysis of data, drafting, and revisions).
ORCID:
Faleh A Khudhair
https://www.orcid.org/0000-0002-3050-5443
HOW TO CITE THIS ARTICLE
Faleh A Khudhair, Shatha H Ali, Khalil A Al-zubeidi. Impact of RAGE Gene rs80096349(C>T), rs1035798(C>T), and rs184003(G>T) Polymorphisms on Non-Response to Aspirin in Iraqi Patients with Coronary Artery Disease. J. Med. Chem. Sci., 2023, 6(7) 1582-1597


